Thirty years ago, The Seattle Times ran a massive Pulitzer Prize-winning series documenting the labyrinthine development of the Boeing 757—the 3 million parts, the 65 miles of wiring, the hundreds of thousands of pages of instructions, the 65,000 “engineering events” behind its design.
“No one really understands the Boeing 757,” wrote Peter Rinearson. “No single person knows how to design or build it. No one human, or even a modest group of humans, could fully fathom its complexities.”
But for outright mystification, a 60-ton jet has nothing on the smallest unit of life, the 1-nanogram cell.
“Cells are so complex,” says Michael Knoblauch, a plant cell biologist. “There is an action of thousands and thousands of substances at a time.”
To fathom it, says Knoblauch, scientists have to break it down into parts, to simplify them until they are understandable. Even if all those parts and individual systems might be comprehensible, conceptualizing their overall workings would be beyond the power of the human brain.
“Compared to a cell,” adds Knoblauch, “an airplane is very simple.”
And visible.
You can see an airplane from ten miles away. Cells are so elusive that even after Robert Hooke described them in 1665, comparing cork cells to the rooms of a monastery, it took almost two more centuries for scientists to theorize the cell was the common unit of development for plants and animals.
Even now, biologists like Knoblauch are flummoxed by a process as seemingly simple as the movement of nutrients from a plant’s leaves to its roots. But they are finding new ways to use two of their profession’s oldest tools: their eyes and the microscope.
In some ways, examining the world with a microscope seems quaint, with so much other science revolving around theory, inference, high-powered data crunching, and modeling, not to mention tools that detect things outside our senses, like the gel electrophoresis that enabled the gene sequencing of the Human Genome Project. The microscope brings with it a simpler seen-with-my-own-eyes satisfaction.
“There’s an excitement from seeing something for the first time every day,” says Chris Davitt, supervisor of WSU’s Franceschi Microscopy and Imaging Center, which deploys half a dozen different microscopes in pursuit of the small. “We’ve all seen E. coli, but then you’re doing an experiment with it and you look and say, wow, that’s what it’s doing or not doing. It’s a pretty incredible job.”
“When you’re working with stuff, oftentimes you can’t see,” says Val Lynch-Holm, the center’s electron microscope specialist. “You’re running gels or you’re trying to make assumptions based on data—this is happening, that is happening. Here you can see. You prep a sample, you look at it—bam—you have instant results. So it’s very rewarding because you’ve done the work and you have a picture right there. And it shows you so much.”
The work also brings with it huge challenges. Like sculptors who must first master welding, the scientists who journey to spaces measured in nanometers—millionths of millimeters—must wrangle photons, electrons, glowing proteins and stains, and samples whose appearance can vary dramatically depending on the microscope at hand and whether the sample is dead or alive. It’s no mere coincidence that Ibn al-Haytham’s work in the field of optics 1,000 years ago led to his now being called the First Scientist. It’s why researchers who master such a craft get to wear two titles: one for their chosen field, like biologist, and a clunky, tongue-tripper for their tool, microscopist.
Michael Knoblauch’s beginnings as a microscopist were inauspicious at best.
At 16, he dropped out of high school.
“I was fed up with just sitting there and learning theory,” he says.
Looking for a more hands-on experience, he signed on as a technical assistant at the Senckenberg Museum in his native Germany, preparing animals for display and making images and graphics for journals and papers. He went on scientific expeditions on the North Sea and near the Alps in southern Germany. He learned to recognize every mouse species in the country.
“It made me learn to see there’s a sense in learning and that it can be fun,” he says. “And I hadn’t had that at school.”
A few years later, while starting his doctorate, he saw an advertisement for someone familiar with the confocal microscope. Capable of making three-dimensional images of cell structures, the tool is one of the most important in the life sciences. But back around 1995, it was so new that people with confocal experience were not to be found.
“They got me,” says Knoblauch.
He has since learned to use a variety of microscopes and become director of the Franceschi center while exploring the inner workings of phloem, the sequence of cells that move nutrients from a plant’s leaves to its roots.
“The big thing, what I’m interested in and still working on, is we don’t understand how plants translocate their nutrients through the phloem,” he says. “What is the driving force for it? … It is as if we didn’t know the heart is driving our circulation. What impact would that have in medicine if you didn’t know that? You wouldn’t know what stroke is. You wouldn’t know what heart attack is. You wouldn’t know all these things.”
Ernst Münch, a German plant physiologist, in 1930 laid out his pressure flow hypothesis to explain the hydrostatic pressure by which fluid moves through the phloem. But nearly 85 years later, scientists still can’t say with confidence that Münch’s hypothesis matches the real world. Knoblauch is hard at work on it, using a variety of tools and protocols that tend to complement each other’s benefits and limitations.
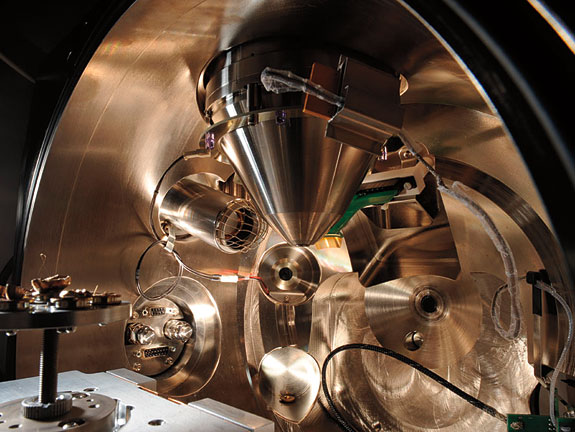
“Because a cell consists of so many different chemicals, like proteins, lipids, all kinds of things, it is difficult to find a good procedure to preserve all of them at once,” he says. He is sitting at a scanning electron microscope, which uses a high-energy beam of electrons to detect a sample’s topography.
This particular sample is from the dehydrated stem of a bean plant. Knoblauch has destroyed its proteins with enzymes and washed out its carbohydrates and lipids with detergent. He cut it with a store-bought razor blade, the sharpest tool short of a diamond knife. He mounted it on a dime-sized plug and coated it with an atomic-thin layer of gold to conduct the microscope’s electrons.
As he starts to look at the sample, the phloem cells, called sieve tubes, appear like holes in a ciabatta. He zooms in, and ever-sharper details loom up like geographic features in Google Earth. At the ends of the cells are microscopic nets called sieve plates, which let liquid move from cell to cell.
“There’s a sieve plate here,” Knoblauch says. “There’s another one. There’s one. There are several inside here, see?”
He continues to zoom; the images continue to stay sharp.
“We are currently at 9,000X,” he says. “We can go about 350,000 with this thing.”
Sieve plates were first seen with a light microscope about 150 years ago, but the diffraction barrier—a resolution limit dictated by a light’s wavelength—meant any two objects within 240 nanometers of each other would appear as one. With this instrument, the resolution is closer to 1 nanometer, the size of about 10 atoms laid end to end.
“This is part of a publication in The Plant Cell in 2010,” says Knoblauch, “and it was the first time that you could see the sieve plates with this detail. This enabled us
to make precise calculations on conductivity of the tubes.”
But there’s more to a sieve tube than a sieve plate, and it happens to have been washed out in preparing the sample. So Knoblauch turns to a transmission electron microscope, which beams electrons through a super-thin sample and can bring all sorts of details into view.
“These are, for example, mitochondria, the organelles that produce the energy, ATP,” says Knoblauch. “Here’s another one. These are membranes, these little things. These are vesicles. This here is a Golgi—you see these layers?”
The ability to see such features helps Knoblauch look at forisomes, fibrous, shape-shifting bodies that plump up and plug sieve plates when a plant is under attack and in danger of springing a high-pressure leak. Knoblauch is one of the world’s leading experts in the forisome, but it’s an elusive feature that managed to hide in plain, albeit microscopic, sight.
“I used the confocal for a very long time and never found the real structure,” he says. That was because he needed to stain what he wanted to see with a fluorescent dye, but since he didn’t know what he was looking for, he didn’t know how to stain it.
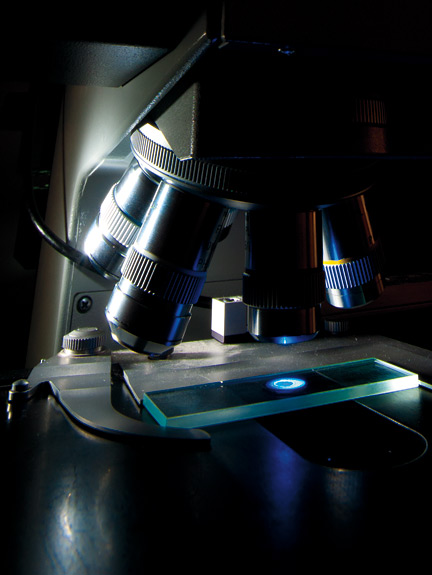
But he knew from old literature that something should be there. He went looking for it on a light microscope, “a standard, old-fashioned, bright field microscope” not far removed from those used back in the day by Robert Hooke and Antonie van Leeuwenhoek, the self-taught Dutchman who discovered sperm cells, bacteria, and protozoa. At first he didn’t see anything and left to have tea with some colleagues. On his return, he saw a toothpick-shaped forisome.
It didn’t look like much, a vague outline, really, but a mere mote to our eyes was a log to Knoblauch.
“If you’re a microscopist, at some point you trust yourself,” he says.
He also suspected the forisome had become visible by changing shape and altering the light properties that suddenly made it visible. So he returned to the confocal microscope and, using different stains, found it in a plumped shape that could prevent sap from leaking out in the event of an injury.
The fact that the forisome took different forms in different microscopes explains how its function eluded investigators for more than a century. It also helps explain why previous microscopists thought they were looking at two different things, if not some random artifact.
“You could never say this is a dynamic reaction from this to this,” Knoblauch says.
For that, a researcher needed to look at the sample live, in vivo. The confocal microscope opened the door to that possibility, giving Knoblauch the chance to see features in their dynamic, natural environment while studying other aspects in greater detail in other types of microscopes.
There’s no orderly sequence of tools in the process, no narrative of microscopes. But often a new instrument like the confocal microscope leads to new protocols and new things to be seen, followed by the arduous process of figuring out what those things are.
“When I see something new, when I know there is something nobody has seen before, that’s one of the biggest things,” say Knoblauch. “Then you have to figure it out. And that often takes years.”
Eric Shelden is looking at a zebrafish. When full-grown, it’s an ordinary, minnow-sized creature that typically has dark, horizontal stripes. This is the descendant of one from the Pullman pet store Barnacle Bill’s. It is an albino—no stripes—and it’s a GloFish®, carrying a gene that will express fluorescent proteins.
It’s also an embryo, under enough magnification that Shelden, an associate professor in the School of Molecular Biosciences, can see nuclei in its cells.
And it’s alive.
“These are living muscle cells in an actual organism in a real tissue,” says Shelden. “These are anesthetized animals, but if they weren’t anesthetized, they could swim. So these are not cells grown on a dish. These are fully differentiated, normal, actual cells in an actual organism in the actual tissue.”
In a 24-hour period, the zebrafish grows from a single cell to having an eye, forebrain, hindbrain, muscle tissue, and a heart.
“You could in principle watch how an eye develops from nothing to a functional eye on this microscope in a day,” says Shelden.
That microscope is the two-photon Leica TCS SP5MP laser scanning confocal microscope. Shelden acquired the device and an ultrafast laser last year with a grant from the M.J. Murdock Charitable Trust, plus matching funds from the dean of research, the Center for Reproductive Biology, and the colleges of Sciences and Veterinary Medicine. It reflects a lot of the changes he’s gone through as a microscopist.
As a kid, he was given a Sears microscope, which he still has, along with a copy of the science classic Microbe Hunters, and he was on his way. Studying biology in grad school, he met Shinya Inoué, one of the first people to connect microscopes to computers, “the grandfather of modern biological microscopy” and a mentor of his graduate advisor.
He also was smitten by what he saw in the cytoplasm of a cell.
“I was just absolutely captivated by that stuff—that each cell in our body has this incredible structure to it,” he says. “It’s not just a little bag of membrane. It’s this really intricate, really beautiful array of structural proteins that gives cells their shape.”
He went on to spend 16 years looking at cell cultures. They’re great for studying, say, the fundamental processes of cell division or how to interfere with cell division in a cancer tumor. But Shelden now speaks like an adherent of seeing the cell in an actual living organism.
“If you want to understand how a neuron behaves physiologically in a low-oxygen environment in the case of something like stroke, well then you have a big problem,” he says, “because neurons don’t act in culture like they act in your brain.”
To start looking at processes deep inside a live animal required several revolutionary developments.
One was the microscope itself. Marvin Minsky, MIT cognitive scientist and son of an ophthalmologist, first conceived of a “confocal scanning microscope” in 1955 to plot neural networks in the brain. But the device needed lasers, computers, and electronic light detectors, all of which happened decades after.
Similarly, two-photon microscopy was conceived of in 1931, but it could not be applied to biology until nearly 60 years later. The technique exposes tissues to less light damage, but it requires bursts of photons so brief—femtoseconds brief—that the first photon in the pulse, traveling at the speed of light, is only a hair’s breadth ahead of the last.
Another important development was green fluorescent protein, which can be used to genetically tag and illuminate proteins in a cell.
“Microscopy was kind of dying for a while with all the genetics and molecular biology and gene mutations,” says Shelden. “All these things were incredibly powerful and didn’t involve fluorescence microscopy. So microscopy was kind of going by the wayside until somebody cloned a gene for a fluorescent protein, and now you can do all kinds of things and you can see the stuff. You can see molecules and proteins… Every day somebody comes up with a new idea to use one of these things.”
Indeed, the week Shelden says this, Time magazine features a green, glow-in-the-dark cat in which the fluorescing protein helps scientists track a gene to fight the feline immunodeficiency virus.
In the zebrafish, Shelden is looking to see if and how a stress-response protein mobilizes in re
sponse to injury or other trauma. The first question was if the protein attaches itself to something in response to injury, like higher temperature.
“The answer is yes it does,” says Shelden, “but only under the right circumstances.
“The next question is what does it attach to? That’s going to be part of what we do next, to repeat this study but look at the specific location of these attachment points.”
If Shelden can figure out how the proteins mobilize to make repairs, that knowledge can be incorporated into therapies, even used to prevent damage before it occurs. Some 10 percent of coronary artery bypass patients have strokes, but it’s possible they could be given activators of stress-response proteins before being operated on.
“We’ll give you a drug that’s going to activate all of these stress responses, and your prognosis for successful surgery is going to go up,” he says.
It’s the kind of research, and result, that microscopists look forward to.
Web exclusive
On the web
WSU Franceschi Microscopy and Imaging Center
Nikon Small World Contest (photomicrography and photovideography)
WSU’s first electron microscope

Built in the 1930s on a shoestring budget with spare and handmade parts, WSU’s original electron microscope was one of the first in the world. (Photo Robert Hubner)
Read more about the history of WSU’s electron microscope.